1. Examples of bacterial cells
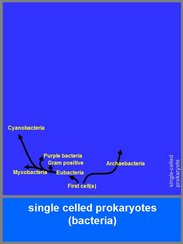
The chances are small that the chemical processes on earth have immediately produced a bacterial cell that was perfect in the sense that it could maintain and reproduce itself. The reason is that many of the first cells that were formed will have died due to errors in the autocatalytic organisation and/or food shortage. On these grounds, it is quite likely that the formation of early cells on earth must have been a rather common process that in many cases led to rapidly dying cells or to lineages of reproducing cells that became extinct after a short period. This implies that the chances on a multiple origin beginning of life on earth are high.
Yet, the number of potential cell types may have been limited from the start, for two reasons. One, the chemical conditions on earth allowed only a limited number of possible autocatalytic sets to be formed. Two, the environment of the newly formed cells selected heavily in favour of certain autocatalytic configurations. At the time when the first cells were formed, the earth had an atmosphere that lacked oxygen. For this reason, the energy to drive chemical reactions had to come from the degradation of abiotic compounds like hydrogen, sulphur, iron, ammonia and nitrite. For example hydrogen can react with carbon dioxide to produce methane, water and energy, or sulphur and hydrogen can fuse to produce hydrogen sulphide and energy. This energy can be used to fuel the production of organic compounds from CO2. It took many millions of years before evolution developed cells that were able to use the radiation energy of the sun: the phototrophic bacteria.
The above shows that an overview of the prokaryotic cell types can be structured best on the basis of the descendance tree of the bacteria, in other words via the bacterial phylogeny. Each major branching in this tree of bacterial evolution represents a major diversification in the autocatalytic set and/or in the construction of the surrounding membrane.
As real archaeology of bacteria is difficult because there are so few detailed fossils of structures with bacterial size, the reconstruction of the bacterial phylogeny requires 'archaeology' using the 'evolutionary clock' of the DNA. Given a certain low mutation rate of the DNA and a common bacterial ancestor, the difference between the DNA of two bacteria species, called their genetic distance, can be used to reconstruct the phylogenetic tree. The longer ago two bacterial species became separate branches on the tree, the more different their DNAs will be. Depending on the selection pressures on the different DNA parts, the molecular clocks may show different rates over time. It may be assumed that a combined picture of descendance trees based on different clocks can yields a good guess at the original diversification processes.
Note that viruses cannot be included in the class of the (prokaryote) cells. The reason is that they do not exist on the basis of a hypercycle-mediating interface. For a large part of the time, separate bits and pieces of the virus float around in the soma of cells showing no individually recognisable autocatalytic hypercycle and no interface. Instead of autocatalysis they could be classified as showing exocatalysis, in the sense that they use the autocatalytic machinery of the host for replication of their viral compounds.
2. Links to interesting sites with information about the bacterial cells
http://www.vermicon.de/images/together/science_figures/tree.jpg (phylogeny graph for bacteria)
http://www.ucmp.berkeley.edu/bacteria/bacteriasy.html (university phylogeny graph)
http://tolweb.org/tree?group=life#TOC2 (phylogeny in tree of life project)
http://www.emunix.emich.edu/~ghannan/biol120/prakaryphylogeny.html (prokaryote phylogeny)
http://www.tulane.edu/~dmsander/WWW/109/Molecular.html (best phylogeny so far!!)
http://www.uwinnipeg.ca/~simmons/1116/16monera.htm (the diversity of prokaryotes)

